 |
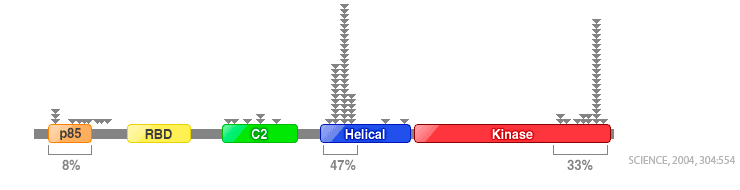 |
PIK3CA mutation activates PI3K (Phosphatidylinositol 3-kinase) protein and PI3K activates several signaling pathways of tyrosine kinase receptors. Activated tyrosine kinase represses the function and excitometabolism of protein which inhibits cell survival and division. These abnormal changes initiate the process of cancer development by activating the protein involved in cell growth. Point mutations in PIK3CA are detected with ca. 80% of frequency in exon 9 (interpreted as helical part of PI3K protein) and exon 20 (interpreted as kinase part), while other types of mutations are also seen in many different locations. PIK3CA mutations are found with 25 to 40 % frequency in various types of cancer like colorectal cancer, gastric cancer, lung cancer, brain cancer, endometrial cancer, ovarian cancer, breast cancer, and more. PNAClamp™ PIK3CA Mutation Detection Kit utilizes modified PCR technology, using optimized PNA probes that tightly bind to wild type DNA templates. This tight binding to the wild type DNA templates result in no amplification of the wild type DNA template in PCR reaction while the mutated genes, especially with SNPs, PCR amplification is processed for multiplication of the mutated DNA sequences.
PNA, (Peptide Nucleic Acid) an artificially DNA analogue, requires less reaction time for hybridization than DNA. It is also highly specific to nucleic acid due to its higher ΔTm value for each single mismatch and highly stable to biological enzyme. These characteristic enable PIK3CA Mutation Detection Kit to detect even a small amount of mutated PIK3CA gene in a short time (ca. 2 hours) with high sensibility (detection limit < 0.5%). Therefore no additional experiment is necessary and only real time PCR reaction with this kit is necessary for detecting mutated genes in PIK3CA.
|
 |
|
Product Name
|
|
Cat. No.
|
|
Size
|
|
PNAClamp™ PIK3CA Mutation Detection Kit
|
|
PNAC-4001 (13 mutations)
|
|
25 tests
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
 |
|
· PNAC-4001(13 mutations)
|
|
No.
|
|
Reagent
|
|
Exon
|
|
Amino Acid Change
|
|
Nucleotide Change
|
|
Cosmic No.
|
|
1
|
|
E542 PNA mix
|
|
9
|
|
p.E542K
|
|
c.1624G>A
|
|
760
|
|
|
|
|
|
|
|
|
|
|
|
p.E542G
|
|
c.1625A>G
|
|
761
|
|
|
|
|
|
|
|
|
|
|
|
p.E542V
|
|
c.1625A>T
|
|
762
|
|
|
|
|
|
|
|
|
|
|
|
|
2
|
|
E545 or Q546 PNA mix
|
|
9
|
|
p.E545K
|
|
c.1633G>A
|
|
763
|
|
|
|
|
|
|
|
|
|
|
|
p.E545G
|
|
c.1634A>G
|
|
764
|
|
|
|
|
|
|
|
|
|
|
|
p.E545D
|
|
c.1635G>T
|
|
765
|
|
|
|
|
|
|
|
|
|
|
|
p.Q546E
|
|
c.1636C>G
|
|
6147
|
|
|
|
|
|
|
|
|
|
|
|
p.Q546K
|
|
c.1636C>A
|
|
766
|
|
|
|
|
|
|
|
|
|
|
|
p.Q546P
|
|
c.1637A>C
|
|
767
|
|
|
|
|
|
|
|
|
|
|
|
p.Q546R
|
|
c.1637A>G
|
|
12459
|
|
|
|
|
|
|
|
|
|
|
|
|
3
|
|
H1047 PNA mix
|
|
20
|
|
p.H1047Y
|
|
c.3139C>T
|
|
774
|
|
|
|
|
|
|
|
|
|
|
|
p.H1047L
|
|
c.3140A>T
|
|
776
|
|
|
|
|
|
|
|
|
|
|
|
p.H1047R
|
|
c.3140A>G
|
|
775
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
 |
 Ready-to-use kit with result available within 3hours. Ready-to-use kit with result available within 3hours.
|
 Compatible Real Time PCR Compatible Real Time PCR
- ABI 7500, ABI 7900, ABI Step one plus, Bio-Rad CFX96, Light Cycler480 (Roche), Rotor-Gene Q.
|
 Compatible Sample Compatible Sample
- Fresh tissue, FFPE tissue.
|
 Test Several Biomarkers in One Plate with same PCR conditions Test Several Biomarkers in One Plate with same PCR conditions
- eg. EGFR+KRAS or EGFR+BRAF+PIK3CA in one plate
|
 High Sensitivity, Specificity with small amount of DNA High Sensitivity, Specificity with small amount of DNA
|
 |
| 1 |
Kwon et al., Frequency of KRAS, BRAF and PIK3CA mutations in advanced colorectal cancers: Comparison of peptide nucleic acid-mediated PCR clamping and direct sequencing in formalin-fixed, paraffin-embedded tissue. Pathol Res Pract 2011;207(12):762-768. |
|
| 2 |
Barbi S. et al., The analysis of PIK3CA mutations in gastric carcinoma and metanalysis of literature suggest that exon-selectivity is a signature of cancer type. J Exp Clin Cancer Res. 2010;29:32. |
|
| 3 |
Di cosimo S. et al., Phosphoinositide 3-kinase mutations in breast cancer: a "good" activating mutation? Clin Cancer Res. 2009;15(16):5017-5019. |
|
| 4 |
Ligresti G. et al., PIK3CA mutations in human solid tumors: role in sensitivity to various therapeutic approaches. Cell Cycle. 2009;8(9):1352-1358. |
|
| 5 |
Kalinsky K. et al., PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res. 2009;15(16):5049-5059. |
|
|